Ph.D. Thesis
The phylogeny of bivalves (Mollusca) consisting of about 20.000 species is still uncertain. Conventional morphology and sequence analyses did not provide conclusive results beyond the four subtaxa Protobranchia, Pteriomorpha, Heterodonta and Schizodonta. This study is concentrated on the folding patterns of slowly evolving 18S rDNA molecules in expectation of finding "deep phylogenetic" information, i.e. pre-Cambrian or Cambrian structural signatures.
Sequences of 18S rDNA with lengths of about 1800 base pairs each, available from Genbank and the European Ribosomal RNA Database, have been studied, covering the four subtaxa. With freely available software, 3D molecular structures were created from the primary sequences und subsequently mapped into 2D patterns. The 2D/3D structures are known to be more conservative than the primary structure (the linear base sequence).
Constructed by a MINIMUM GIBBS FREE ENERGY (MFE) PRINCIPLE, these foldings show high variability, on average at about 40 strand sites. By contrast, "NEAR-NATURAL" STRUCTURES folded with a structural constraint show much less variability and much higher stability of the characters to be classified.
The phylogenetic analysis of the MFE STRUCTURES revealed that despite of some phylogenetic information the matrix included too much 'noise' to result in statistic support of deep nodes. In the first preliminary phylograms derived, PHYLOGENETIC GROUPS WERE CLEARLY DISCERNIBLE BUT ALWAYS INTERSPERSED WITH RUNAWAYS for unclear reasons. A clear result for bivalve phylogeny could not be found.
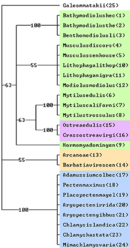
A "NEAR-NATURAL" FOLDING shows much less variability in its structure. More than 80% of it is conservative leaving only few and small clusters for morphological analysis. Even more bases than in MFE structures are bound in double bindings thereby restricting the development of new morphological characters through point mutation. A preliminary analysis of a character matrix of 24 pteriomorph species clearly supports previously discerned families – most taxa remain monophyletic, with the exception of paraphyletic Mytiloidea. A clear resolution within the families was due to too few characters not to be expected. All the same it promises good results in deeper nodes.
18S rDNA folded with structural information in RNAViz with highlighted molecular morphological characters Simple parsimony analyses of the constraint character matrix of pteriomorph bivalves
Supported by the German Science Foundation (DFG), project HA2598/9-1,2 in SPP 1174 "Deep Metazoan Phylogeny"